[ad_1]
Common genetic variants may underlie autism spectrum disorder and schizophrenia across human populations, according to a study appearing September 11th in the journal Cell Reports. In line with previous studies in Caucasians, the researchers found that Japanese individuals with autism spectrum disorder and schizophrenia have overlapping copy number variations (CNVs) — inter-individual variations in the number of copies of a particular gene.
“The strength of our study is the systematic head-to-head comparison of pathogenic CNVs and biological pathways between autism spectrum disorder and schizophrenia,” says senior study author Norio Ozaki of Nagoya University Graduate School of Medicine. “Previous studies in Caucasian populations found overlap in pathogenic CNVs between the two disorders, but their analyses were limited to a small number of genes and CNV loci.”
Autism spectrum disorder and schizophrenia have complex inheritance patterns, with multiple genetic and environmental factors influencing disease risk. Available evidence points to genetic overlap between the two clinically distinct disorders. For example, they tend to co-occur at a higher rate than would be expected in the general population, and a large epidemiological study showed that a family history of schizophrenia in first-degree relatives is a risk factor for autism spectrum disorder. In particular, previous studies have revealed that these two disorders are associated with an increased burden of CNVs, and that rare CNVs in specific loci are shared risk factors for both disorders.
However, the majority of the previous CNV studies were carried out in Caucasian populations, limiting the generalization of pathogenic CNVs and relevant biological pathways. Studies in populations other than Caucasians may also provide additional biological insights into the disorders. Moreover, the clinical features of patients with pathogenic CNVs have not been fully examined in non-Caucasian populations. Until now, no studies have directly compared pathogenic CNVs and biological pathways between autism spectrum disorder and schizophrenia in non-Caucasian populations.
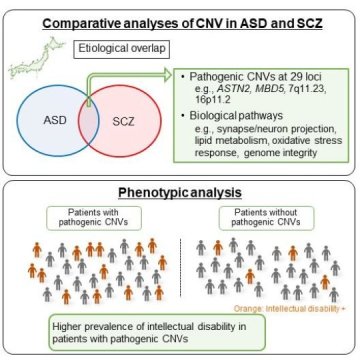
To address this gap in knowledge, Ozaki and his team performed comparative CNV analyses of 1,108 cases of autism spectrum disorder, 2,458 individuals with schizophrenia, and 2,095 controls in a Japanese population using a high-resolution technique called array comparative genomic hybridization. They confirmed an increased genome-wide burden of rare CNVs in autism spectrum disorder and schizophrenia and observed an overlap in pathogenic CNVs between the two disorders. Pathogenic CNVs, including those at 29 loci common to both disorders, were found in about 8% of the two types of patients, which was significantly higher than in controls.
“Genetic overlap has been suggested in epidemiological and molecular genetic studies,” says first author Itaru Kushima of Nagoya University Graduate School of Medicine. “In line with this, our systematic and comprehensive investigation confirmed a significant overlap of pathogenic CNVs between autism spectrum disorder and schizophrenia in a Japanese population.”
Additional analysis revealed that both disorders are associated overlapping biological pathways involved in the oxidative stress response, lipid metabolism, and genomic integrity. The researchers also identified 12 CNV loci potentially associated with these disorders in a Japanese population. Moreover, intellectual disability was strongly associated with pathogenic CNVs in both patient groups. “The identification of shared pathways and disease-relevant genes provides biological insights into autism spectrum disorder and schizophrenia,” Ozaki says.
Moving forward, the researchers plan to investigate CNVs in bipolar disorder and examine the overlap of pathogenic CNVs and biological pathways among the three disorders. In the near future, they will also conduct studies using CNV-based animal models, as well as induced pluripotent stem cells (iPSCs) derived from patients with pathogenic CNVs.
Story Source:
Materials provided by Cell Press. Note: Content may be edited for style and length.
[ad_2]