[ad_1]
One method cells have to reliably curb the amount of specific proteins at any desired time is the control on mRNA degradation. As the mRNA’s tail plays a role in this process, biologists at the Center for RNA Research, within the Institute for Basic Science (IBS, South Korea), have identified how mixed tails — made of different nucleotides — protect mRNA from degradation for longer. Published in Science, these findings could bring new insights to our understanding of gene regulation in healthy and diseased states.
mRNA is a delicate molecule with an important function; it brings genetic information contained in the DNA out of the nucleus to produce proteins. When a gene is transcribed into messenger RNA (mRNA), the mRNA is equipped with a tail, that acts like a shield against pre-mature degradation.
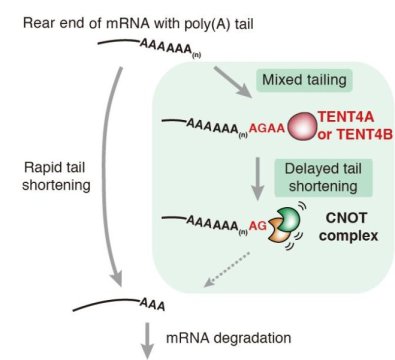
Until recently, mRNA tails were thought to be merely a chain of hundreds of adenosine (A) monophosphate nucleotides, and was thus commonly called the poly(A) tail. Specific enzymes lengthen and shorten the tail by adding and pruning A’s from its end: poly(A) polymerases add a chain of around 200 A’s; and deadenylases, such as the CNOT complex, trim down the poly(A) tails over time starting from the A’s at the tip of the tail.
In 2014, IBS researchers discovered that the mRNA tail is not limited to A’s. They developed a high-throughput sequencing method, TAIL-seq, to accurately measure the length of poly(A) tails at a genome-wide scale. They found out that other nucleotides other than A’s, like guanosine (G), uridine (U) and cytosine (C), decorated the tail end. Mixed tails were reported in an evolutionarily broad range of species, including: humans, mice, frogs, and fish, adding further importance to this topic.
In this current study, IBS researchers found out that some of the enzymes that insert A’s to the tail, are able to add also G’s, U’s, and C’s, creating a mixed tail. In particular, nucleotidyltransferase TENT4A/B intermittently adds G’s when extending the mRNA tail. Interestingly, in cells G’s are located mainly at the very end of the tail, or at the next to last position. This can be explained by the fact that the enzymes that prune the poly(A) tails stall when they encounter a G, rather than an A, at the end of the tail. In other words, the research team found out that a G addition may slow down the trimming of the tail, thereby protecting it.
“Non-adenosines added by TENT4A/B have a considerable stalling effect. Just one is enough to counteract poly(A) trimming enzymes and stabilize the mRNA,” explains Young-suk Lee, one of the authors of the study. The mRNAs with mixed tails were indeed reduced in cells lacking TENT4A/B.
“The mRNA tail has been considered a pure stretch of A’s with little informational content, except for its length. However, this study proves that even tails of the same length can have different composition, and a mixed tail degrades more slowly than the pure poly(A) tail,” explains KIM Narry, corresponding author of the study and director of the Center. “Through this study, we can understand the life history of complex mRNA by revealing a new kind of mRNA protection mechanism that was not known before.”
In the future, the team would like to expand their research to various biological systems, and to understand how misbehaving tailing mechanisms can lead to various diseases. In addition, they would like to develop a RNA-based gene therapy method that utilizes the mixed tailing effect in cells to enhance mRNA stability.
Story Source:
Materials provided by Institute for Basic Science. Note: Content may be edited for style and length.
[ad_2]