Immune checkpoint inhibitors have revolutionized cancer care. The therapy works by preventing tumors from shutting down the immune response, which in turn allows T cells to kill cancer cells. Established checkpoint inhibitors target the proteins PD-1 and CTLA-4 and are used to treat a variety of solid tumor types, including melanoma and lung cancer. However, the U.S. Food and Drug Administration recently approved a new immune checkpoint inhibitor targeting the protein LAG3. This anti-LAG3 antibody, called relatlimab, was administered in combination with the anti-PD-1 antibody nivolumab to treat advanced melanoma.
Despite this therapeutic breakthrough, little has been known about the structure of the LAG3 protein. In the absence of a three-dimensional structure, LAG3-based drugs must be designed “in the dark” using inefficient screening methods. A team of Moffitt Cancer Center researchers has become the first in the world to visualize the molecular structure of the LAG3 protein. In a new article published in Nature Immunology, they describe the crystal structure of LAG3 and how it interacts with molecules produced by cancer cells.
“When I started my lab at Moffitt, I noticed a growing interest in LAG3 as an immunotherapy target. I was surprised at how little we knew about the LAG3 structure and its molecular mechanism, despite about 30 years of literature highlighting its role in the immune system,” said Vince Luca, Ph.D., assistant member of the Drug Discovery Department.
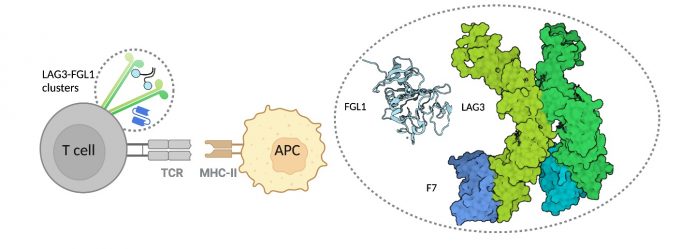
Luca and his team used X-ray crystallography to “see” the structure of the LAG3 protein at nearly atomic resolution. The researchers also mapped out the regions of LAG3 that bound to signaling molecules called FGL1 and MHCII and two different anti-LAG3 antibodies. From this information, they were able to determine which antibody binding sites were ideal to inhibit LAG3 activity.
Through their investigations, the researchers discovered how structural interactions of LAG3 and FGL1 inhibit T cell function. They found that binding of the two molecules causes LAG3 to cluster on the surface of T cells, which they hypothesize may contribute to the inhibitory activity of LAG3 by blocking the T cells from properly recognizing tumor cells.
These combined data reveal several important insights into the three-dimensional structure of LAG3 and how it interacts with other molecules, which may lead to better targeted therapeutic approaches in the future.
“Collectively, our structural, epitope mapping and functional studies provide an improved framework for understanding LAG3 molecular function. In the future, additional structures of LAG3 bound to ligands and antibodies will refine our knowledge of the LAG3 signaling axis to illuminate how extracellular binding events fine-tune LAG3-mediated changes in T cell activity. In turn, such structural insights should guide the development of maximally effective LAG3-based immunotherapies,” said Luca.