[ad_1]
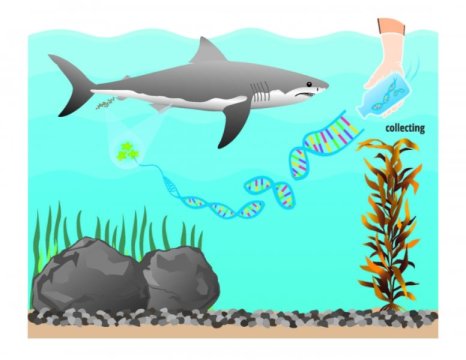
A white shark’s acute sense of smell is legendary, allowing it to detect a potential meal several miles away — and giving pause to those of us who work and play in the ocean.
But now humans can sniff them out as well, thanks to a collaboration between researchers from UC Santa Barbara and the U.S. Geological Survey, with colleagues from California State University Long Beach (CSULB) and Central Michigan University (CMU). Through developments in environmental DNA (eDNA), scientists — and soon, perhaps, any curious individual — can determine if white sharks have been nearby.
“One of the goals of this research is for a lifeguard to be able to walk down to the shore, scoop up some water, shake it and see if white sharks are around,” said Kevin Lafferty, a USGS ecologist and researcher with UCSB’s Marine Science Institute (MSI). He is lead author of the new paper “Detecting southern California’s white sharks with environmental DNA,” published in the journal Frontiers in Marine Science.
The team’s results add to a growing body of evidence that white sharks, which had been declining in numbers due to overfishing, have for the last several years been experiencing a comeback along the California coast.
According to CSULB professor, white shark expert and study co-author Chris Lowe, the resurgence is due to the success of state and federal protections from fishing, recovery of marine mammal populations and better fisheries management.
“However, white shark population recovery has co-occurred during a period when more people than ever before are using the coastal ocean for recreation, ultimately increasing the likelihood of interactions,” he said. “While sightings of juvenile white sharks have risen considerably along California over the last eight years, there has been no dramatic increase in shark bites on people.”
Environmental DNA is genetic material collected from the environment, as opposed to within a living organism. Things animals may leave behind — such as mucus, feces or shed skin — contain their genetic signatures, which can be parsed out and identified through genetic sequencing. Scientists can extract and amplify specific genes within the DNA fragments found in water samples, and determine if the DNA contained in those samples is from a specific species.
In Carpinteria, down the coast from UC Santa Barbara, Lowe had been acoustic- and satellite-tracking tagged juvenile sharks at one of several summer/fall nurseries along southern California. Lafferty and Lowe wondered if these sharks were leaving a detectable eDNA plume. Lafferty had little success using eDNA to sample for sharks until eDNA expert and MSI researcher Chris Jerde shared a new protocol he developed with CMU’s Andrew Mahon.
“Ten years ago we started working on eDNA,” said Jerde, who is a co-author of the paper. “The advances in technology since then have dramatically improved the reliability, portability and widespread application of the method.”
Using a new species-specific genetic analysis called digital droplet PCR, CMU biology professor and study co-author Mahon designed specific genetic markers from white shark tissue sent to him by Lowe. Mahon’s student, Kasey Benesh, analyzed the water samples and controls blindly. Samples near the shark aggregation matched the genes in the white shark tissue, whereas water samples a mile away did not, confirming that a water sample could sniff out white sharks.
Because eDNA can drift with currents, and sharks can swim long distances in the time it takes eDNA to degrade, the new approach only gives a rough idea about where sharks actually are at a particular moment. Still, “Chris Lowe can now add eDNA to his new white shark monitoring program, which includes real-time acoustic tracking and drone flights,” Lafferty said.
For surfers, ocean swimmers and beachgoers, the increase in white shark population may be a cause for concern. Although white sharks don’t feed on humans (and juveniles favor rays and other fish), they have certainly been known to bite out of defensiveness, curiosity or mistaken identity, causing grave or lethal injuries. Environmental DNA monitoring could give lifeguards and other people responsible for public safety clues as to when to be extra vigilant, and also help marine biologists understand how well white sharks are recovering in response to protection.
“We can now sample eDNA along the coast to make better maps and seasons for white sharks,” Lafferty said, “And if we can do it for white sharks, we can do it for other marine species, too.”
Added Lowe, “We can use eDNA not only to determine whether white sharks have been present at a beach, but also to determine if their favorite food is there is well, such as stingrays. Once we are able to better refine and calibrate the methods, another goal will be to integrate eDNA technology into autonomous surface vehicles that can be programmed to move along the coast sampling water and send data into the cloud, along with text alerts to local lifeguards, of the presence of white sharks at a particular location. This technology holds great promise for future, near real-time monitoring.”
The result of the scientists’ study proves that eDNA sequencing has become a powerful tool for tracking the general movement of single species, as well as for monitoring the biodiversity of a region in real time. Initially a tool to detect the presence of invasive species, such as Asian carp and bullfrogs, eDNA has also been used to detect threatened and endangered species that are often elusive, such as arroyo toads, California red-legged frogs and tidewater gobies.
“New advances in eDNA are allowing for not just a single species to be detected, but instead the DNA for the water sample is screened for all fish species or all amphibian species,” Jerde said. “The same technology used to decode the human genome is now used to sequence all the DNA in a water sample. From this we can monitor fish stocks, measure the presence or absence of rare species, and better connect how climate change and pollution are impacting biodiversity.”
This study was made possible with support from UCSB’s Marine Biodiversity Observation Network.
[ad_2]