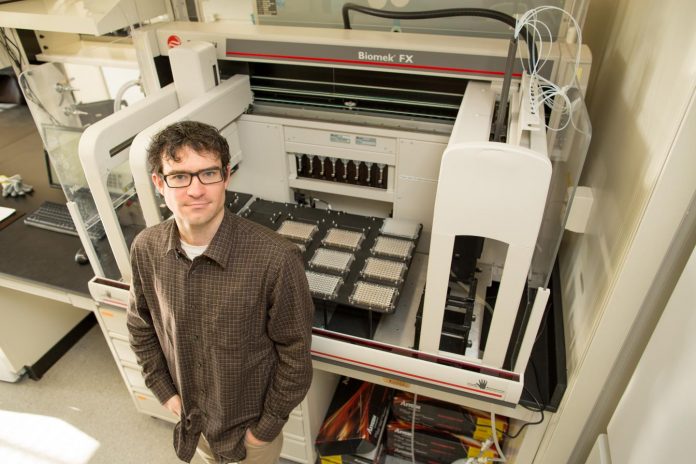
Photo: Gregory Lang, an associate professor in the Department of Biological Sciences at Lehigh University, in his lab.
view more
Credit Image: Douglas Benedict/Academic Image
If B is better than A, and C is better than B, it follows by the transitive property that C is better than A. And, yet, this is not always the case. Every kid is familiar with the Rock-Paper-Scissors game–the epitome of nontransitivity in which there is no clear hierarchy among the three choices, despite each two-way interaction having a clear winner: Paper beats Rock, Scissors beats Paper, and Rock beats Scissors.
Evolution may be teeming with nontransitive interactions as well. While natural selection – the process by which organisms better adapted to their environments are more likely to survive and pass on their genes – can be observed over shorter time intervals, there is still debate about whether fitness gains accumulate over long evolutionary time scales. In other words, one might expect that successive adaptive events (like the two-way interactions of Rock-Paper-Scissors) would translate into a cumulative increase in fitness, resulting in the very latest generation always being more fit than its all of its genealogical ancestors. However, this turns out to not be true in every case.
The evolutionary process, then, includes what are known as nontransitive interactions, sometimes producing organisms that are less fit than its ancestors. Experimental demonstrations of such nontransitivity, however, have been lacking.
Until now. A group of scientists at Lehigh University led by Gregory Lang, associate professor in the Department of Biological Sciences, has recently provided empirical evidence that evolution can be nontransitive. Lang and his team identify a nontransitive evolutionary sequence through a 1,000-generation yeast evolution experiment. In the experiment, an evolved clone outcompetes a recent ancestor but loses in direct competition with a distant ancestor.
The nontransitivity in this case arose as a result of multilevel selection that involved adaptive changes in both the yeast nuclear genome and the genome of an intracellular RNA virus. The results, which provide experimental evidence that the continuous action of selection can give rise to organisms that are less fit compared to a distant ancestor, are described in an article published in eLife Journal today called “Adaptive evolution of nontransitive fitness in yeast” (DOI: 10.7554/eLife.62238).
This study confronts two common misconceptions about evolution, according to Lang. The first, he says, is that evolution is a linear “march of progress” where each organism along a line of descent is more fit than all those that came before it.
Lang and his colleagues set out to determine how nontransitivity arose along a particular line of genealogical descent. In their 1,000-generation yeast experiment, nontransitivity arose due to adaptation in the yeast nuclear genome combined with the stepwise deterioration of an intracellular virus. Initially the population produced a virally encoded toxin and was immune to the toxin. As the population adapted, it fixed the beneficial nuclear mutations as well as mutations within the intracellular viral population that resulted in loss of toxin production. Over time the more beneficial nuclear mutations fix, and selection in the viral population resulted in a loss of toxin immunity – since the toxin was no longer produced. When placed in competition against its distant ancestor, the 1,000-generation evolved population lost due to the toxin produced by the ancestor.
“Another misconception is that there is a single locus of selection,” says Lang. “Multilevel selection–as its name implies–states that selection can act simultaneously on multiple levels of biological organization.”
In the context of this experiment, multilevel selection was common, says Lang. “Selection acts across multiple levels of biological organization, from genes within a cell to individuals within a population. Selection at one level can impact fitness at another.
“In fact, when we expanded our study of host-virus genome evolution to additional populations, we found that nearly half of the approximately 140 populations we studied experienced multilevel selection, fixing adaptive mutations in both the nuclear and viral genomes,” he adds.
“Laboratory evolution experiments have proven highly effective for studying evolutionary principles, yet this work is the first to document a non-transitive interaction and provide a mechanistic explanation,” says co-author Sean W. Buskirk, an assistant professor at West Chester University who collaborated on the research when a postdoctoral student in Lang’s lab. “Ultimately, the presence of a virus in the ancestor drastically impacts how the evolved yeast populations compete and interact with one another.”
The work of co-author Alecia B. Rokes, at the time an undergraduate biology major at Lehigh, focused on competing two intracellular viruses inside yeast cells in what she terms her very own “virus fight club.”
“I worked on competing two viruses within the yeast cells to see if either virus variant had an advantage over the other, thus leading to higher frequency and one virus outcompeting the other,” says Rokes, now a graduate student in microbiology at the University of Pittsburgh. “It was amazing to be part of the process of elimination, persistence, and pure curiosity that went into figuring out what was actually going on in these populations.”
By showing that nontransitive interactions can arise along a line of genealogical succession, the team’s work has broad implications for the scientific community’s understanding of evolutionary processes.
“It resolves what evolutionary biologist Stephen Jay Gould referred to as ‘the paradox of the first tier,’ which is the failure to identify broad patterns of progress over long evolutionary time scales, despite clear evidence of selection acting over successive short time intervals,” says Lang. “In addition, it calls into doubt whether true fitness maxima exist and, more broadly, it implies that directionality and progress in evolution may be illusory.”
###
TDnews (tunisiesoir.com)