[ad_1]
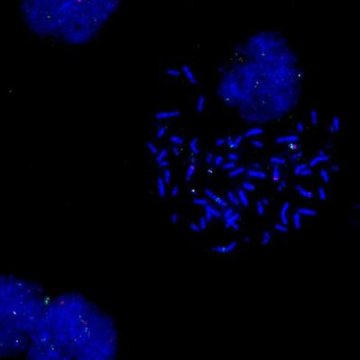
Cancer cells often have mutations in their DNA that can give scientists clues about how the cancer started or which treatment may be most effective. Finding these mutations can be difficult, but a new method may offer more complete, comprehensive results.
A team of researchers has developed a new framework that can combine three existing methods of finding these large mutations — called structural variants — into a single, more complete picture. The new method — published in Nature Genetics on September 10, 2018 — could help researchers find new structural variations within cancer cell DNA and learn more about how those cancers begin.
“We were able to design and use this computational framework to connect the three methods together, to get the most comprehensive view of the genome,” says Feng Yue, assistant professor of biochemistry and molecular biology at the Pennsylvania State University and one of the paper’s corresponding authors. “Each method by itself can only review a portion of the structural variations, but when you integrate the results of the three different methods, you can get the most comprehensive view of the cancer genome.”
Structural variants are large mutations in DNA that can result in cancer-causing genes to be turned on. For example, certain types of brain cancer, such as the types that afflicted U.S. Senators John McCain and Edward Kennedy, can be caused by structural variants that amplify certain cancer-causing genes. In some cancers, knowing a patient has this abnormality helps doctors decide on the best treatment plan.
The researchers say finding these structural variants is important for several reasons.
“Many of the structural variants that are found in human cancers do not appear to directly impact a gene,” says Jesse Dixon, a Helmsley-Salk Fellow at the Salk Institute and one of the paper’s first and corresponding authors. “Instead, many structural variants appear in noncoding portions of the genome, what people have historically referred to as junk DNA, and it can be a bit of a mystery as to why these may be contributing to cancer.”
Adds Yue, “If you are a cancer patient, knowing about the structural variants that lead to the cancer can help us understand why you got sick and possibly which treatment could be best.”
The researchers used three existing methods for finding structural variants: optical mapping, high-throughput chromosome conformation capture (known as Hi-C), and whole genome sequencing, which has been used to discover the majority of structural variants that are already known.
Using their new method, the researchers were able to find structural variations for over 30 types of cancer cells. The team of scientists also used the same methods to begin to learn why certain classes of structural variants may be contributing to cancers.
What the researchers observed is that some structural variants appear to affect regulatory gene “switches” in noncoding sequences of DNA. Defective switches prevent appropriate turning on or off specific genes and this can contribute to cancer. They also used the Hi-C method to explore how structural variations can affect 3D genome structure — how DNA folds itself within the cell.
“Cells are small, but their DNA is very long — laid out in a line, all the DNA from one cell would be more than two meters long,” says Job Dekker, professor and co-director of the program in Systems Biology at University of Massachusetts, investigator of the Howard Hughes Medical Institute and a corresponding author of the paper. “That’s why DNA needs to fold in intricate ways. We have found that genomic alterations in cancer cells can lead to differences in how the genome folds and this can lead to cases where genes become turned on or off by the wrong regulatory switches.”
The team of scientists was able to discover that structural variants impact genome folding in cancer cells, and that these changes may be contributing to the cancer.
“One of discoveries we had made in the past is that our genome is folded up into distinct structures, almost like little neighborhoods,” Dixon says. “It appears as though some structural variants cause changes to these neighborhoods, such that a cancer causing gene is moved from a neighborhood where the gene is kept quiet into one where the gene becomes activated.”
The scientists suggest that this work could lead to a better ability to predict which structural variants may contribute to cancer, and which genes they may be targeting.
The National Institute of Health, Howard Hughes Medical Institute, European Research Council, Cancer Research UK, Wellcome Trust helped support this work.
Other authors included: Jie Xu, Fan Song, Yanli Wang, Lijun Zhang, Hongbo Yang, Tingting Liu, Sriranga Iyyanki, Lin An, James R. Broach and Ross C. Hardison of Pennsylvania State University; Vishnu Dileep, Takayo Sasaki, Juan Carlos Rivera-Mulia and David M. Gilbert of Florida State University; Ye Zhan, Hakan Ozadam, Bryan R. Lajoie and Job Dekker of University of Massachusetts; Victoria T. Le of Salk; Galip Gürkan Yardimci and William Stafford Noble of University of Washington; Abhijit Chakraborty and Ferhat Ay of La Jolla Institute for Allergy and Immunology; Darrin V. Bann and Christopher Pool of Milton S. Hershey Medical Center; Royden Clark of Penn State College of Medicine; Rajinder Kaul, Michael Buckley, Kristen Lee, Morgan Diegel and John A. Stamatoyannopoulos of Altius Institute for Biomedical Sciences; Dubravka Pezic and Suzana Hadjur of University College London; and Christina Ernst and Duncan T. Odom of Cambridge University.
[ad_2]