[ad_1]
In the last six years, a tool called CRISPR-Cas9 has transformed genetic research, allowing scientists to snip and edit DNA strands at precise locations like a pair of tiny scissors.
But sometimes, it takes more than scissors to do the job. Now, a collaborative international team has unveiled a new CRISPR-based tool that acts more like a shredder, able to wipe out long stretches of DNA in human cells with programmable targeting.
Writing in Molecular Cell, they describe how they succeeded in getting a different kind of CRISPR-Cas system called Type I CRISPR-Cas3 to work as long-range DNA editing tool in human cells for the first time.
The resulting tool provides a way to target and delete much longer expanses of DNA than current Cas9 tools can. That power could be put to work in genetic research to understand the underpinnings of disease — and potentially in treatment for diseases tied to long stretches of DNA.
Yan Zhang, Ph.D., the University of Michigan scientist who led the research, explains that the new tool harnesses a different type of CRISPR system than the Cas9-containing ones widely used . Both are borrowed from bacteria, where they normally function to find and weed out invading DNA.
A different type of CRISPR
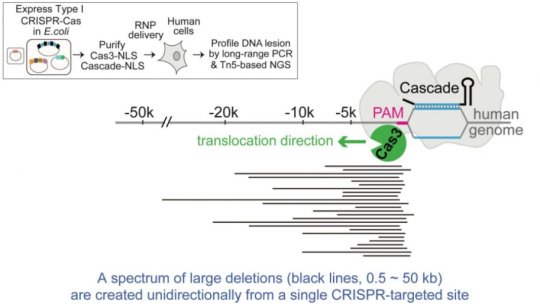
The new tool uses a Type I CRISPR, which is much more common in bacteria than the Type II variety that includes Cas9. Type I CRISPR has never been used in any eukaryotic cells, and employs a riboprotein complex known as Cascade for seeking its target and an enzyme called Cas3 for shredding DNA.
The challenging protein optimization and purification side of the work was done in the laboratory of Cornell University professor Ailong Ke, Ph.D., a co-corresponding author of the paper. Cornell graduate student Adam Dolan and U-M senior research specialist Zhonggang Hou, Ph.D. are the paper’s first authors.
The research involved a long shot by Zhang, who has studied bacterial CRISPR-Cas9 and developed tools for editing genetic material in human cells, and Ke, who has studied Type I CRISPR using structural and biochemical approaches.
They set out to try to deliver the bacterial CRISPR components as proteins into both human embryonic stem cells and another type of cell called HAP1. With one CRISPR guide, the team succeeded in deleting portions of targeted DNA ranging from a few hundred base pairs to 100 kilobases.
A motorized shredder
Zhang calls the Cascade-Cas3 system a “DNA shredder with a motor” because it can move along a DNA genome for a certain distance, breaking up the genetic material as it goes.
“Cas9 is a molecular scissor that goes where you want it and snips once,” says Zhang, an assistant professor of biological chemistry at the U-M Medical School. “But Cas3 goes where you want it, travels along the chromosome, and makes a spectrum of deletions tens of kilobases long. This could make it a powerful screening tool to determine what large areas of DNA are most important for a particular disease.”
This could be especially useful when scientists are studying the “non-coding” long stretches of DNA that don’t contain the code for a particular protein; the “shredder” technique could allow them to demolish a sequence of long stretches and see what happens.
In addition, the ability of Cas3 to travel on the chromosome over a long distance can’t be done with any current Cas9 technique. So a “nuclease dead” version of Cas3 that can travel along DNA but lacks the shredder function might provide a powerful delivery platform for long-range epigenome engineering.
Scientific challenges
Part of the research effort involved figuring out how to get the human stem cells to reveal if any DNA had been deleted, since most of the stem cell “reporter” lines developed for research involving CRISPR-Cas9 are not sensitive enough if the shredding activity is low. Sara Howden, Ph.D. of the University of Melbourne, Australia, worked on building a sensitive, dual-reporter cell line.
Another challenge for the team was to figure out what the shredder had shredded after the deed was done, using next-generation DNA sequencing and going beyond existing methods to look at the small edits that Cas9 produces.
Hou developed a transposase-based profiling method, and Peter Freddolino, Ph.D., an assistant professor of biological chemistry and computational medicine and bioinformatics at U-M, built custom informatics pipelines to analyze the deep sequencing results.
Ke also notes in the paper that because the guide RNA sequence needed to tell the “shredder” where to go is longer than the sequences used in CRISPR-Cas9 systems, and also because target searching and degradation are two well-separated steps, the new method is potentially better controlled. It may be less likely to make mistaken cuts where they’re not wanted.
This new tool and its derivatives could be useful for therapeutic purposes, Zhang speculates, though such use is years in the future.
Some CRISPR-Cas9-based therapeutic uses have been reported — including a controversial editing out of a receptor gene that enables HIV to enter cells in the embryos of two babies reportedly born in China. But worries about CRISPR-Cas9 making unintended edits in normal areas of human patients’ DNA have also surfaced. Further work will be needed to see if the “shredder” approach avoids this issue.
U-M and Cornell have filed for a joint patent on this new tool. The research was funded by the National Institutes of Health (GM128637, GM118174, GM102543 and GM117268) and by Zhang’s funding from the Medical School’s Biological Sciences Scholars Program.
[ad_2]